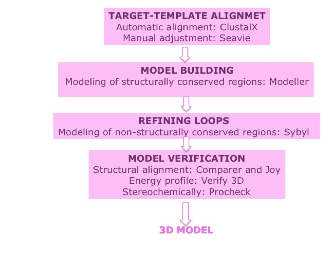
The homology modeling consists in several steps. The firs step is to obtain a correct sequence alignment of the target sequences with the homologous template, using program as CLUSTALW followed by a manual adjustment of the multiple alignment sequence with the program SEAVIEW. From the best alignment, 3D models containing all non-hydrogen atoms are obtained automatically using the method implemented in MODELLER. The structurally variable regions (SVRs) as loops are built with the standard procedure in MODELLER and afterwards they are refined. After the refinement process the models are validated using the VERIFY ,PROCHECK and COMPARER programs.